FISH
(Restore this version)
Modified: 18 March 2018, 11:42 AM User: Luca Torello Pianale →
FISH
Fluorescent in situ hybridization (FISH) is a molecular cytogenetic technique used to detect the presence of a specific DNA sequence on a chromosome.
The technique is based on the use of a probe: a short DNA sequence (10-25 nucleotides), specifically designed for the target sequence. To allow the detection of a wanted genic locus the probe is fluorescent-labelled (direct labelling) or can be rendered fluorescent in a subsequent phase (indirect labelling).
The first protocol step is the attachment of either interphasic or metaphasic chromosomes on a solid surface (glass); then the sample and probe solution is denatured and incubation starts. During the incubation period (hours, depending on the protocol), target DNA and probe interact and hybridize. To avoid background, unbound/partially bound probes are washed away, by repeated washes, after incubation.
The detection step can be different: in case of direct detection the probe is already fluo-marked and the sample can be directly observed at fluorescence microscope. On the contrary, in the indirect labelling it is necessary to add an enzymatic or immunological detection system, which will render the probe fluorescent. This method needs more time but allows to amplify the signal and get a better detection.
FISH is a versatile technique: can be useful to detect a gene locus on a chromosome, chromosomal abnormalities and rearrangements. Nowadays several connected techniques have been developed starting form the “classical” FISH, which is focused on chromosomal loci.
(Giada Cipollina)
Immuno-FISH
As the name suggests, this is a combination of two techniques, the standard FISH, either on flattened chromosome preparations (2-D FISH) or on three-dimensionally preserved nuclei (3-D FISH), and the other indirect or direct immunofluorescence. Immunofluorescence permits the detection of nuclear proteins (modified histones, histone variants and modifiers, transcription machinery and factors, nuclear sub-compartments, etc), so that both DNA and proteins can be analyzed on the same sample. Numerous methods involving a variety of fixation and permeabilization techniques can be used for immunofluorescence applications, and the choice depends on cell type, epitope, and antibody being used.
Some protocols establish that FISH procedure comes first, but not all antigens will be preserved after the various steps in the protocols, so it is possible to apply the primary or primary and secondary antibodies and proceed with a fixation step prior to the FISH procedure: in this way, the detected epitope will be preserved.
The combination of these techniques is often used to investigate co-localization of genomic regions with proteinaceous entities within interphase nuclei such as nucleoli or promyelocytic leukemia (PML) bodies. It has helped to position chromosomes in interphase nuclei.
(Fabiola Varese)
RNA-FISH
As we already said, FISH is a very powerful method to detect optically specific nucleic acid sequences (both DNA and RNA sequences) in fixed but otherwise intact cells. DNA FISH is useful to understand the number of gene copies, the location of the gene in the nucleus when it is turned on/off, DNA repair and replication studies. However, RNA-FISH is a technique that has become a very powerful tool to study gene expression and RNA biology. In fact, RNA-FISH is a quantifiable method that can determine the localization of single mRNA molecules in the cell, their number, their transcription and degradation rates and more.
There are two categories of RNA-FISH: those that use some form of signal amplification, and those that rely on direct detection of a signal.
Direct Detection
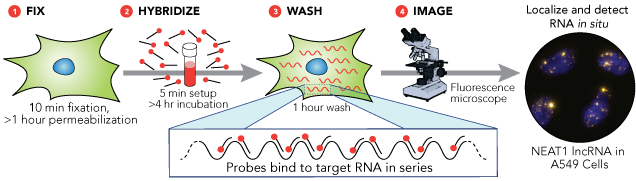
Direct detection involves labeling the probes themselves with fluorophores (the probes must have enough fluorescence to be detectable above background autofluorescence). One technique is to use a set of short single-stranded DNA oligonucleotides complementary to various regions of the target RNA, each labelled with one or more fluorescent moieties. The binding of multiple probes enables the localisation of the target RNA via fluorescence microscopy as a fluorescent spot. The advantage of this approach is that the off-target binding of a single oligonucleotide in the probe pool will either be undetectable or readily distinguishable to the much brighter spots corresponding to the true RNA, thus reducing the chances of false positives. False negatives are similarly unlikely, for even if a single probe out of the pool fails to bind, the rest are likely to bind.
Signal Amplification

In order to circumvent the limitation of low signals from the relatively small numbers of fluorescent molecules targeted to mRNA in these direct detection methods (A), researchers have also developed a large variety of methods to amplify signals from individual molecules. Some of these are relatively simple extensions of the direct detection methods, such as detection of the probe by fluorescently labelled antibodies targeting specific haptens (small molecule that reacts with specific antibody but is not immunogenic by itself) incorporated in large numbers into an RNA probe (B). Others involve targeting nucleic acid probes with a single or few haptens with antibodies conjugated to enzymes; those enzymes in turn will act upon a substrate in such a manner as to create a fluorescent product that will become covalently linked to surrounding molecules (C).
Another technique is the RNAScope Assay (Figure Below). Here, we produce two unlabeled tandem probes, containing a short complementary region (18-25 bases), a spacer sequence and a 14-base tail sequence (rapresented as “Z” in the image). After hybridization with the target probes, comes a second hybridization step with a pre-amplifier probe. This is a long probe that contains a complementary sequence to the 28 bases of the two target probes tails (14+14). Therefore, only when the two target tails hybridize one next to the other the pre-amplifier will hybridize. The pre-amplifier contains 20 binding sites for an amplifier probe, which in turn contains 20 binding sites for the labelled probe. Thus, for each target probe pair, we get 20×20=400 labelled probes. Such method has the advantage of labeling targets to such a degree that the signals are easily visible even by eye, precluding the need for expensive optical setups. In addition, they are able to reliably detect short RNA molecules such as miRNA. However, such methods are somewhat prone to lower detection efficiencies owing to the large number of steps in such protocols, each of which has some probability of failure.

Figure References:
1. https://biosearchassets.blob.core.windows.net/assets/standard/PAGE/2239/stellaris-method.png
2. https://ars.els-cdn.com/content/image/1-s2.0-S1046202316300123-gr1.jpg
3. https://greenfluorescentblog.wordpress.com/2013/02/21/rnascope-a-novel-fish-in-the-sea/
(Luca
Torello)
others....